New Zealand's economy is dependent on agriculture, a sector that is highly sensitive to climate change. This makes it critical to develop analysis capabilities to assess its impact and investigate possible mitigation and adaptation options. That analysis can be done with tools such as agricultural systems models. In simple terms, it involves creating a model to quantify how a specific crop behaves under certain conditions then simulating altering a few variables to see how that behavior changes. Some of the software available to do this includes CropSyst from Washington State University and the Agricultural Production Systems Simulator (APSIM) from the Commonwealth Scientific and Industrial Research Organization (CSIRO) in Australia.
Historically, these models have been used primarily for small area (point-based) simulations where all the variables are well known. For large area studies (landscape scale, e.g., a whole region or national level), the soil and climate data need to be upscaled or downscaled to the resolution of interest, which means increasing uncertainty. There are two major reasons for this: 1) it is hard to create and/or obtain access to high-resolution, geo-referenced, gridded datasets; and 2) the most common installation of crop modeling software is in an end user's desktop or workstation that's usually running one of the supported versions of Microsoft Windows (system modelers tend to prefer the GUI capabilities of the tools to prepare and run simulations, which are then restricted to the computational power of the hardware used).
New Zealand has several Crown Research Institutes that provide scientific research across many different areas of importance to the country's economy, including Landcare Research, the National Institute of Water and Atmospheric Research (NIWA), and the New Zealand Institute for Plant & Food Research. In a joint project, these organizations contributed datasets related to the country's soil, terrain, climate, and crop models. We wanted to create an analysis framework that uses APSIM to run enough simulations to cover relevant time-scales for climate change questions (>100 years' worth of climate change data) across all of New Zealand at a spatial resolution of approximately 25km2. We're talking several million simulations, each one taking at least 10 minutes to complete on a single CPU core. If we were to use a standard desktop, it would probably have been faster to just wait outside and see what happens.
Enter HPC
High-performance computing (HPC) is the use of parallel processing for running programs efficiently, reliably, and quickly. Typically this means making use of batch processing across multiple hosts, with each individual process dealing with just a little bit of data, using a job scheduler to orchestrate them.
Parallel computing can mean either distributed computing, where each processing thread needs to communicate with others between tasks (especially intermediate results), or it can be "embarrassingly parallel" where there is no such need. When dealing with the latter, the overall performance grows linearly the more capacity there is available.
Crop modeling is, luckily, an embarrassingly parallel problem: it does not matter how much data or how many variables you have, each variable that changes means one full simulation that needs to run. And because simulations are independent from each other, you can run as many simulations as you have CPUs.
Solve for dependency hell
APSIM is a complex piece of software. Its codebase is comprised of modules that have been written in multiple different programming languages and tightly integrated over the past three decades. The application achieves portability between the Windows and GNU/Linux operating systems by leveraging the Mono Project framework, but the number of external dependencies and workarounds that are required to run it in a Linux environment make the implementation non-trivial.
The build and install documentation is scarce, and the instructions that do exist target Ubuntu Desktop editions. Several required dependencies are undocumented, and the build process sometimes relies on the binfmt_misc kernel module to allow direct execution of .exe files linked to the Mono libraries (instead of calling mono file.exe), but it does so inconsistently (this has since been fixed upstream). To add to the confusion, some .exe files are Mono assemblies, and some are native (libc) binaries (this is done to avoid differences in the names of the executables between operating system platforms). Finally, Linux builds are created on-demand "in-house" by the developers, but there are no publicly accessible automated builds due to lack of interest from external users.
All of this may work within a single organization, but it makes APSIM challenging to adopt in other environments. HPC clusters tend to standardize on one Linux distribution (e.g., Red Hat Enterprise Linux, CentOS, Ubuntu, etc.) and job schedulers (e.g., PBS, HTCondor, Torque, SGE, Platform LSF, SLURM, etc.) and can implement disparate storage and network architectures, network configurations, user authentication and authorization policies, etc. As such, what software is available, what versions, and how they are integrated are highly environment-specific. Projects like OpenHPC aim to provide some sanity to this situation, but the reality is that most HPC clusters are bespoke in nature, tailored to the needs of the organization.
A simple way to work around these issues is to introduce containerization technologies. This should not come as a surprise (it's in the title of this article, after all). Containers permit creating a standalone, self-sufficient artifact that can be run without changes in any environment that supports running them. But containers also provide additional advantages from a "reproducible research" perspective: Software containers can be created in a reproducible way, and once created, the resulting container images are both portable and immutable.
-
Reproducibility: Once a container definition file is written following best practices (for instance, making sure that the software versions installed are explicitly defined), the same resulting container image can be created in a deterministic fashion.
-
Portability: When an administrator creates a container image, they can compile, install, and configure all the software that will be required and include any external dependencies or libraries needed to run them, all the way down the stack to the Linux distribution itself. During this process, there is no need to target the execution environment for anything other than the hardware. Once created, a container image can be distributed as a standalone artifact. This cleanly separates the build and install stages of a particular software from the runtime stage when that software is executed.
-
Immutability: After it's built, a container image is immutable. That is, it is not possible to change its contents and persist them without creating a new image.
These properties enable capturing the exact state of the software stack used during the processing and distributing it alongside the raw data to replicate the analysis in a different environment, even when the Linux distribution used in that environment does not match the distribution used inside the container image.
Docker
While operating-system-level virtualization is not a new technology, it was primarily because of Docker that it became increasingly popular. Docker provides a way to develop, deploy, and run software containers in a simple fashion.
The first iteration of an APSIM container image was implemented in Docker, replicating the build environment partially documented by the developers. This was done as a proof of concept on the feasibility of containerizing and running the application. A second iteration introduced multi-stage builds: a method of creating container images that allows separating the build phase from the installation phase. This separation is important because it reduces the final size of the resulting container images, which will not include any dependencies that are required only during build time. Docker containers are not particularly suitable for multi-tenant HPC environments. There are three primary things to consider:
1. Data ownership
Container images do not typically store the configuration needed to integrate with enterprise authentication directories (e.g., Active Directory, LDAP, etc.) because this would reduce portability. Instead, user information is usually hardcoded explicitly in the image directly (and when it's not, root is used by default). When the container starts, the contained process will run with this hardcoded identity (and remember, root is used by default). The result is that the output data created by the containerized process is owned by a user that potentially only exists inside the container image. NOT by the user who started the container (also, did I mention that root is used by default?).
A possible workaround for this problem is to override the runtime user when the container starts (using the docker run -u… flag). But this introduces added complexity for the user, who must now learn about user identities (UIDs), POSIX ownership and permissions, the correct syntax for the docker run command, as well as find the correct values for their UID, group identifier (GID), and any additional groups they may need. All of this for someone who just wants to get some science done.
It is also worth noting that this method will not work every time. Not all applications are happy running as an arbitrary user or a user not present in the system's database (e.g., /etc/passwd file). These are edge cases, but they exist.
2. Access to persistent storage
Container images include only the files needed for the application to run. They typically do not include the input or raw data to be processed by the application. By default, when a container image is instantiated (i.e., when the container is started), the filesystem presented to the containerized application will show only those files and directories present in the container image. To access the input or raw data, the end user must explicitly map the desired mount points from the host server to paths within the filesystem in the container (typically using bind mounts). With Docker, these "volume mounts" are impossible to pre-configure globally, and the mapping must be done on a per-container basis when the containers are started. This not only increases the complexity of the commands needed to run an application, but it also introduces another undesired effect…
3. Compute host security
The ability to start a process as an arbitrary user and the ability to map arbitrary files or directories from the host server into the filesystem of a running container are two of several powerful capabilities that Docker provides to operators. But they are possible because, in the security model adopted by Docker, the daemon that runs the containers must be started on the host with root privileges. In consequence, end users that have access to the Docker daemon end up having the equivalent of root access to the host. This introduces security concerns since it violates the Principle of Least Privilege. Malicious actors can perform actions that exceed the scope of their initial authorization, but end users may also inadvertently corrupt or destroy data, even without malicious intent.
A possible solution to this problem is to implement user namespaces. But in practice, these are cumbersome to maintain, particularly in corporate environments where user identities are centralized in enterprise directories.
Singularity
To tackle these problems, the third iteration of APSIM containers was implemented using Singularity. Released in 2016, Singularity Community is an open source container platform designed specifically for scientific and HPC environments. "A user inside a Singularity container is the same user as outside the container" is one of Singularity's defining characteristics. It allows an end user to run a command inside of a container image as him or herself. Conversely, it does not allow impersonating other users when starting a container.
Another advantage of Singularity's approach is the way container images are stored on disk. With Docker, container images are stored in multiple separate "layers," which the Docker daemon needs to overlay and flatten during the container's runtime. When multiple container images reuse the same layer, only one copy of that layer is needed to re-create the runtime container's filesystem. This results in more efficient use of storage, but it does add a bit of complexity when it comes to distributing and inspecting container images, so Docker provides special commands to do so. With Singularity, the entire execution environment is contained within a single, executable file. This introduces duplication when multiple images have similar contents, but it makes the distribution of those images trivial since it can now be done with traditional file transfer methods, protocols, and tools.
The Docker container recipe files (i.e., the Dockerfile and related assets) can be used to re-create the container image as it was built for the project. Singularity allows importing and running Docker containers natively, so the same files can be used for both engines.
A day in the life
To illustrate the above with a practical example, let's put you in the shoes of a computational scientist. So not to single out anyone in particular, imagine that you want to use ToolA, which processes input files and creates output with statistics about them. Before asking the sysadmin to help you out, you decide to test the tool on your local desktop to see if works.
ToolA has a simple syntax. It's a single binary that takes one or more filenames as command line arguments and accepts a -o {json|yaml} flag to alter how the results are formatted. The outputs are stored in the same path as the input files are. For example:
$ ./ToolA file1 file2
$ ls
file1 file1.out file2 file2.out ToolAYou have several thousand files to process, but even though ToolA uses multi-threading to process files independently, you don't have a thousand CPU cores in this machine. You must use your cluster's job scheduler. The simplest way to do this at scale is to launch as many jobs as files you need to process, using one CPU thread each. You test the new approach:
$ export PATH=$(pwd):${PATH}
$ cd ~/input/files/to/process/samples
$ ls -l | wc -l
38
$ # we will set this to the actual qsub command when we run in the cluster
$ qsub=""
$ for myfiles in *; do $qsub ToolA $myfiles; done
...
$ ls -l | wc -l
75Excellent. Time to bug the sysadmin and get ToolA installed in the cluster.
It turns out that ToolA is easy to install in Ubuntu Bionic because it is already in the repos, but a nightmare to compile in CentOS 7, which our HPC cluster uses. So the sysadmin decides to create a Docker container image and push it to the company's registry. He also adds you to the docker group after begging you not to misbehave.
You look up the syntax of the Docker commands and decide to do a few test runs before submitting thousands of jobs that could potentially fail.
$ cd ~/input/files/to/process/samples
$ rm -f *.out
$ ls -l | wc -l
38
$ docker run -d registry.example.com/ToolA:latest file1
e61d12292d69556eabe2a44c16cbd27486b2527e2ce4f95438e504afb7b02810
$ ls -l | wc -l
38
$ ls *out
$Ah, of course, you forgot to mount the files. Let's try again.
$ docker run -d -v $(pwd):/mnt registry.example.com/ToolA:latest /mnt/file1
653e785339099e374b57ae3dac5996a98e5e4f393ee0e4adbb795a3935060acb
$ ls -l | wc -l
38
$ ls *out
$
$ docker logs 653e785339
ToolA: /mnt/file1: Permission deniedYou ask the sysadmin for help, and he tells you that SELinux is blocking the process from accessing the files and that you're missing a flag in your docker run. You don't know what SELinux is, but you remember it mentioned somewhere in the docs, so you look it up and try again:
$ docker run -d -v $(pwd):/mnt:z registry.example.com/ToolA:latest /mnt/file1
8ebfcbcb31bea0696e0a7c38881ae7ea95fa501519c9623e1846d8185972dc3b
$ ls *out
$
$ docker logs 8ebfcbcb31
ToolA: /mnt/file1: Permission deniedYou go back to the sysadmin, who tells you that the container uses myuser with UID 1000 by default, but your files are readable only to you, and your UID is different. So you do what you know is bad practice, but you're fed up: you run chmod 777 file1 before trying again. You're also getting tired of having to copy and paste hashes, so you add another flag to your docker run:
$ docker run -d --name=test -v $(pwd):/mnt:z registry.example.com/ToolA:latest /mnt/file1
0b61185ef4a78dce988bb30d87e86fafd1a7bbfb2d5aea2b6a583d7ffbceca16
$ ls *out
$
$ docker logs test
ToolA: cannot create regular file '/mnt/file1.out': Permission deniedAlas, at least this time you get a different error. Progress! Your friendly sysadmin tells you that the process in the container won't have write permissions on your directory because the identities don't match, and you need more flags on your command line.
$ docker run -d -u $(id -u):$(id -g) --name=test -v $(pwd):/mnt:z registry.example.com/ToolA:latest /mnt/file1
docker: Error response from daemon: Conflict. The container name "/test" is already in use by container "0b61185ef4a78dce988bb30d87e86fafd1a7bbfb2d5aea2b6a583d7ffbceca16". You have to remove (or rename) that container to be able to reuse that name.
See 'docker run --help'.
$ docker rm test
$ docker run -d -u $(id -u):$(id -g) --name=test -v $(pwd):/mnt:z registry.example.com/ToolA:latest /mnt/file1
06d5b3d52e1167cde50c2e704d3190ba4b03f6854672cd3ca91043ad23c1fe09
$ ls *out
file1.out
$Success! Now we just need to wrap our command with the one used by the job scheduler and wrap all of that again with our for loop.
$ cd ~/input/files/to/process
$ ls -l | wc -l
934752984
$ for myfiles in *; do qsub -q short_jobs -N "toola_${myfiles}" docker run -d -u $(id -u):$(id -g) --name="toola_${myfiles}" -v $(pwd):/mnt:z registry.example.com/ToolA:latest /mnt/${myfiles}; doneNow that was a bit clunky, wasn't it? Let's look at how using Singularity simplifies it.
$ cd ~
$ singularity pull --name ToolA.simg docker://registry.example.com/ToolA:latest
$ ls
input ToolA.simg
$ ./ToolA.simg
Usage: ToolA [-o {json|yaml}] <file1> [file2...fileN]
$ cd ~/input/files/to/process
$ for myfiles in *; do qsub -q short_jobs -N "toola_${myfiles}" ~/ToolA.simg ${myfiles}; doneNeed I say more?
This works because, by default, Singularity containers run as the user that started them. There are no background daemons, so privilege escalation is not allowed. Singularity also bind-mounts a few directories by default ($PWD, $HOME, /tmp, /proc, /sys, and /dev). An administrator can configure additional ones that are also mounted by default on a global (i.e., host) basis, and the end user can (optionally) also bind arbitrary ones at runtime. Of course, standard Unix permissions apply, so this still doesn't allow unrestricted access to host files.
But what about climate change?
Oh! Of course. Back on topic. We decided to break down the bulk of simulations that we need to run on a per-project basis. Each project can then focus on a specific crop, a specific geographical area, or different crop management techniques. After all of the simulations for a specific project are completed, they are collated into a MariaDB database and visualized using an RStudio Shiny web app.
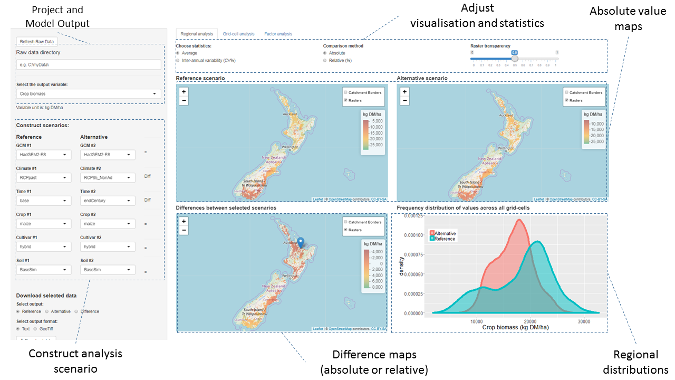
Prototype Shiny app screenshot shows a nationwide run of climate change's impact on maize silage comparing current and end-of-century scenarios.
The app allows us to compare two different scenarios (reference vs. alternative) that the user can construct by choosing from a combination of variables related to the climate (including the current climate and the climate-change projections for mid-century and end of the century), the soil, and specific management techniques (like irrigation or fertilizer use). The results are displayed as raster values or differences (averages, or coefficients of variation of results per pixel) and their distribution across the area of interest.
The screenshot above shows an example of a prototype nationwide run across "arable lands" where we compare the silage maize biomass for a baseline (1985-2005) vs. future climate change (2085-2100) for the most extreme emissions scenario. In this example, we do not take into account any changes in management techniques, such as adapting sowing dates. We see that most negative effects on yield in the Southern Hemisphere occur in northern areas, while the extreme south shows positive responses. Of course, we would recommend (and you would expect) that farmers start adapting to warm temperatures starting earlier in the year and react accordingly (e.g., sowing earlier, which would reduce the negative impacts and enhance the positive ones).
Next steps
With the framework in place, all that remains is the heavy lifting. Run ALL the simulations! Of course, that is easier said than done. Our in-house cluster is a shared resource where we must compete for capacity with several other projects and teams.
Additional work is planned to further generalize how we distribute jobs across compute resources so we can leverage capacity wherever we can get it (including the public cloud if the project receives sufficient additional funding). This would mean becoming job scheduler-agnostic and solve the data gravity problem.
Work is also underway to further refine the UI and UX aspects of the web application until we are comfortable it can be published to policymakers and other interested parties.
If you are interested in our work from a scientific point of view, please contact me and I will put you in touch with the project leader. For all other inquiries, you can also contact me and I will do my best to help.
Eric Burgueño will present Using containers to analyse the impact of climate change and soil on New Zealand crops at linux.conf.au, January 21-25 in Christchurch, New Zealand.






3 Comments